Gene Function Search is a page where users can search for genes from various datasets available in GDC (Genomic Database for Cannabis). Users can search for genes from annotated whole genome assemblies, by location, name, accession or function..
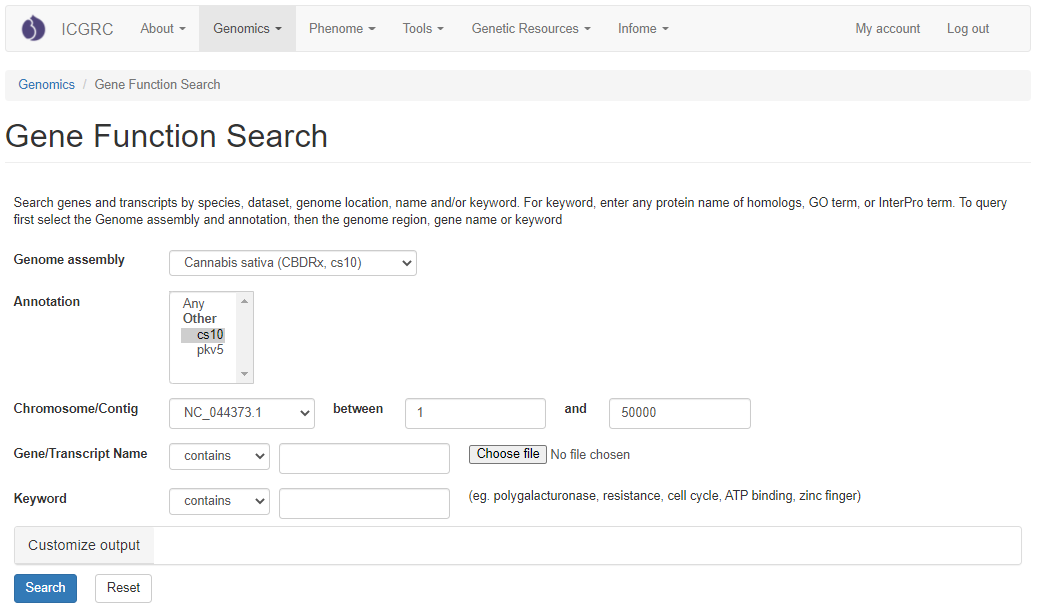
Please Note: All the search categories below, except the file upload, can be combined.
1. Genome assembly
Use these drop-down menus to limit results to genes from a specific genome assembly. Once an assembly is chosen, chromosome/contigs are dynamically populated in the dropdown.
2. Annotation
Use this drop-down menu to limit the results to genes from a specific annotation. Go to 'Description of Sequence Dataset in GDC' for more details. Multiple options can be selected by holding down the "Ctrl" key.
3. Chromosome/Contig
Users can limit their results of predicted genes by their genome location. When a genome assembly is chosen in the drop-down menu next to 'Genome assembly', the corresponding chromosome or scaffold names are dynamically displayed in the 'Chromosome/Contig' drop-down menu. Choose any option and then type in the position in bp in the text boxes.
4. Gene Name
Users can search genes by name for an exact match, contains, starts with or ends with the input, by selecting the desired option from the drop-down menu. The search is case-insensitive. Example gene names are TPS23, LOC... Gene names in a file, separated by a new line, can be uploaded to do a batch search.
3. Keyword
Users can limit their result by associated functional terms. Predicted genes from whole genome assembly and transcripts have been annotated with some of the followings: homology to genes of closely related or plant model species, InterPro protein domains, GO terms, KEGG pathway and ortholog terms. Users can enter any protein name (eg. polygalacturonase), KEGG term/EC number (eg. resistance, EC:1.4.1.3), GO term (eg. cell cycle, ATP binding), or InterPro term (eg. zinc finger) in the text box to limit the results with the entries that are associated with the functional annotation terms.
