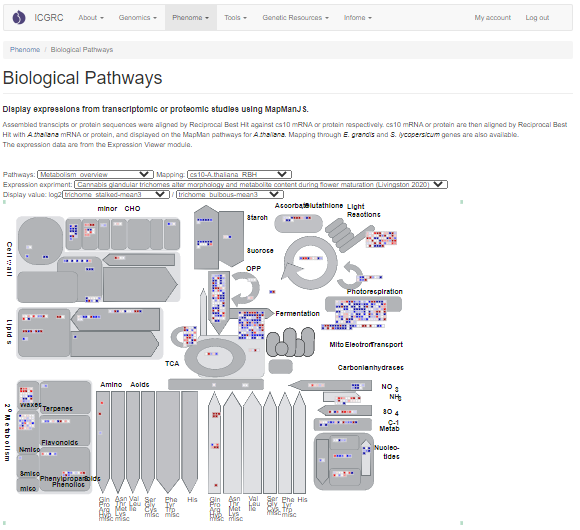
The expression data can also be viewed over biological pathways. It is an instance of MapManJS, a web-based implementation of MapMan for visualizing expression data on metabolic pathways and biological processes.
1. Pathways - select pathway to display
2. Mapping - select gene mapping to use. Mappings are precomputed using blastp reciprocal best hit between proteins, and blastn reciprocal best hit between mRNAs if no protein match is found. Transcripts from transcriptome experiments are matched with cs10, which in turn are matched with either the A. thaliana, E. grandis (eucalyptus) or L. lycopersicum (tomato) mRNAs used by MapMan.
3. Expression experiment - source of expression data/publication
4. Display value - value to display, blue means >0 and red means <0. For experiments with absolute expression values, the user can opt to display either the absolute value or the log2 ratio between two samples by selecting the sample in the denominator. Some experiments only have log2 fold change data available, for these the denominator option should be kept to 1.